Kettle Sour Bacterial Cell Counting
Craft breweries use bacterial cultures in the kettle sour process for sour beers. The kettle sour trend is increasing because of growing demand for new and unique beer flavors. This method of sour beer production is preferred because the wort can be sterilized later to lower the risk of bacterial contamination in the brewery. Calculating cell counts and determining proper pitch rates are crucial for brewing success. Yeast cell counting and pitch rates for beer fermentation are well established in the industry. This is not the case for bacterial cell counting and pitch rates which are equally important for the final beer quality.
Last October, our QC team presented a poster on Lactic Acid Bacteria (LAB) pitch rates at the national Master Brewers Association of the Americas (MBAA) conference in Seattle. Below is a brief synopsis of the project that was presented!
Experiment
The purpose of the experiment was analyzing the effects of pitch rates of LAB on kettle sour to determine if higher initial concentrations of LAB can expedite the souring process. The expedited souring allows brewers to have a faster turn-around time, allowing less time for contamination to occur during the souring process and potentially produce a kettle soured wort within the same brew day or next morning.
The three LAB strains tested were Lactobacillus brevis, Lactobacillus delbrueckii, and Pediococcus pentosaceus from BSI’s culture bank. A 10°P no-hop wort was pitched at 5 x 106 cells/ml, 1 x 107 cells/ml, and 1.5 x 107 cells/ml. At various intervals, the cell count was performed using a Cellometer assay developed for counting bacteria. The stain used for fluorescing the bacteria was SYTO-BC. The pH was measured using a pH probe (Ross).
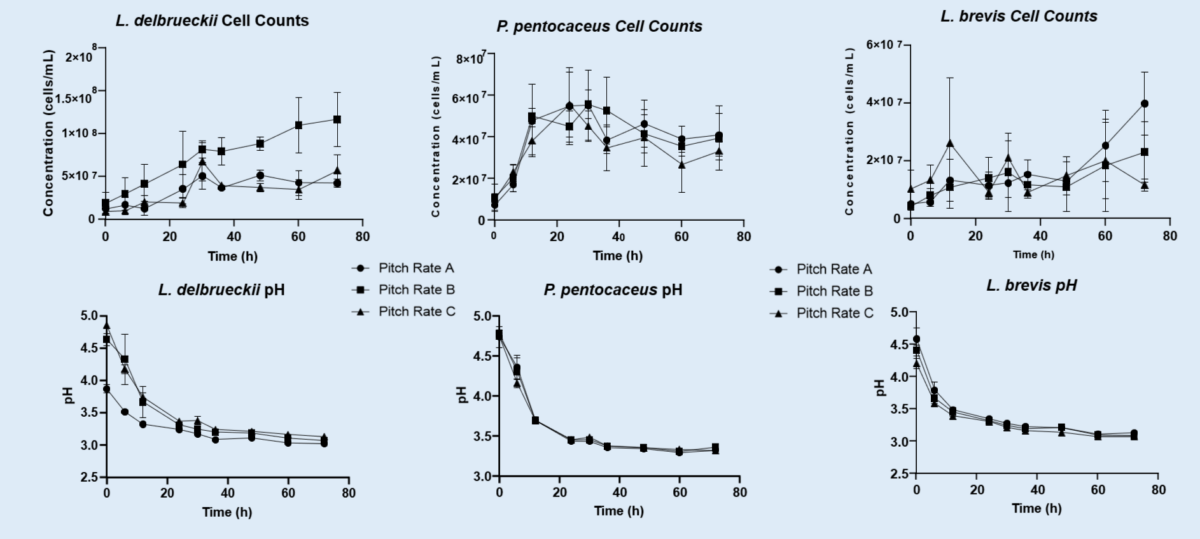 Between all three pitch rates tested, terminal pH and peak cell density were achieved around 24-hour period. Some pitch rates had slight differences in pH at hour 6. But with further time the differences were less noticeable. It was determined that the higher pitch rates with more drastic differences should be studied to see if a quicker kettle souring process is achievable.
Between all three pitch rates tested, terminal pH and peak cell density were achieved around 24-hour period. Some pitch rates had slight differences in pH at hour 6. But with further time the differences were less noticeable. It was determined that the higher pitch rates with more drastic differences should be studied to see if a quicker kettle souring process is achievable.
Cellometer Bacteria Cell Count
Counting bacteria via Cellometer is a relatively quick process once your Cellometer and working stain solution are fine tuned to your specific needs. The working solution needs to be no more than 24 hours old and stored properly. It is important to note that the bacteria cell counts are only accurate between concentrations of 2 x 106 cells/ml and 2.5 x 107 cells/ml. The stained and diluted solution should rest for 10 mins in a dark environment to ensure proper fluorescence. After the stained sample is loaded into the SD025 counting chamber, the slide needs to settle for 20-30 seconds. The slide ends should be covered with clear tape to slow the “Brownian motion” of the solution.
Focusing the Cellometer is another critical step. Without proper focus the samples will give inaccurate and unoptimized cell counts. The cells should be barely visible, the focus is between the cells appearing as a dark speck and a light halo. The user can preview the fluorescent view, and if the stained cells pop a bright green, then the focus is ideal. The first or second cell counts are going to have the strongest fluorescence, so be mindful and precise.
This experiment was run by Nic Chapman, Kory Davis, Patrick Munn, Brandon Munoz, and Nava Thapa of BSI. The Cellometer assay was developed by Revvity, and the data from this experiment was analyzed with assistance from Dr. Leo Chan and Mackenzie Pierce of Revvity.

